 No, the cell is not equivalent to a test tube...
No, the cell is not equivalent to a test tube... 
 No, the cell is not equivalent to a test tube...
No, the cell is not equivalent to a test tube... 
 |
 |
 |
Structural studies continuously uncovers the importance of organization in biological systems at different levels, ranging from molecules to whole organisms. However, many studies of biological function and regulation are performed with isolated systems, in which essential features of the original organization have been altered or lost. This approach forces the experimenter to treat the systems as independent entities for which a set of assumptions apply, but do not necessarily reflect their natural operating conditions. Other approaches involving the macroscopic study of the same systems in their native (non isolated) state yield global information which is not readily amenable to interpretation in terms of their mechanism of action and control. The comparison of the intrinsic properties of isolated systems with their effective behavior in situ generally indicate a most significant contribution of the native organization.
Although immense progress has been independently achieved in each of these "main stream" aspects, they tend to diverge into separate specialized fields, and no comprehensive approach is presently available to readily integrate the resulting accumulated information. The "missing links" involve two major groups of modulating factors including
The considerations above apply to virtually all biological systems. While broadly recognized, they are seldom applied or effectively taken into account in functional studies of processes beyond the strictly molecular level. The knowledge of the mode of interaction between systems among which matter or energy is transfered (e.g., chemical reactants, oxido-reduction equivalents, osmotic or mechanical work, signals, etc.), would not only improve significantly the basic understanding of biological function, but also enable the identification and eventually the design of better systems for use in technological applications.
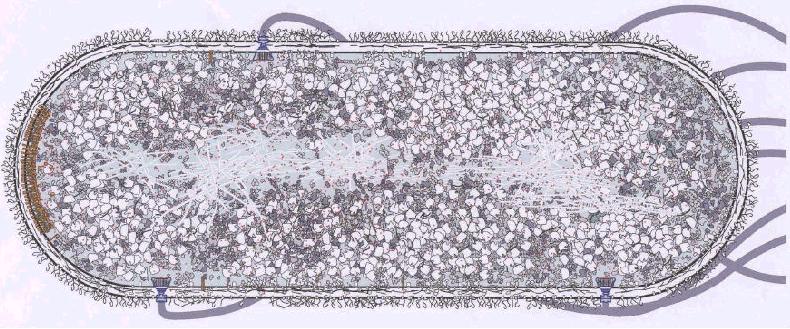
Biological organization ("cellular socialism") must lead to various functional consequences, often arbitrarily segregated into qualitative and quantitative aspects. Both relate to observed phenomena that cannot be explained solely on the basis of the behavior of chemical (or catalyzed) reactions in a homogeneous solution, in which the reactants are assumed to be randomly distributed and equally accessible to each other ("molecular democracy").
In the intracellular milieu, the intermediate reactants of a metabolic sequence may be transferred directly between consecutive enzymic components without equilibrating with the ambient medium. This arises from either the existence of a stable complex consisting of the enzymes in the sequence, or their pair-wise transient association, during which the product of one enzyme is directly transferred to the next one. This phenomenon is reflected experimentally by the lack of availability of the intermediate in solution (e.g., with respect to isotopic dilution, or to an exogenous enzymic trap). It effectively results in an alteration of the expected kinetic properties of the whole pathway, when a simple homogeneous behavior is assumed. Such channeling of intermediates is carried out through molecular recognition between the catalytic components, which involve specific interactions at their respective surfaces. Alternatively, the association of an enzyme with a non-substrate ligand (small molecule, enzyme, or structural element) may result in the allosteric modulation of its catalytic activity.
The reason for the need of such complex mechanisms, too often left aside for philosophers, is probably to promote flexible means for control of cellular processes. Indeed, the possibility of selective channeling of intermediates enables preferential routing of metabolic fluxes through branched pathways, in the crowded cellular environment. In contrast, a freely diffusible intermediate, serving as a substrate to different enzymes randomly distributed in an homogeneous solution, would be subject to simple competition and its fate predetermined by the fixed concentrations of the binding sites.
Limitation by diffusion of the kinetic mechanism of an enzyme acting in solution is considered as the ultimate criterion for "catalytic perfection". It is a fact that most enzymes considered as soluble did not evolve to reach this exalted status. It is likely that the selective constrains on efficiency relate also to the interactive aspects of the various systems rather than to their performance as independent entities only. Accordingly, optimal cellular function has been attained through the simultaneous evolution of favorable interactions between the systems. Finally, it should be borne in mind that the organization in biological systems is not restricted to space; the components may be dynamically rearranged or interact differently as a response to changes in the ambient conditions according to metabolic needs, and thus operate differently in time.
As mentioned in section 1, no established approach is able to address directly the question of functional interaction between related systems in situ, on the basis of their properties observed in vitro after their isolation. One can nevertheless single out proper means to demonstrate and study these interactions.
|
|
|
|